Welcome to Current Exchange
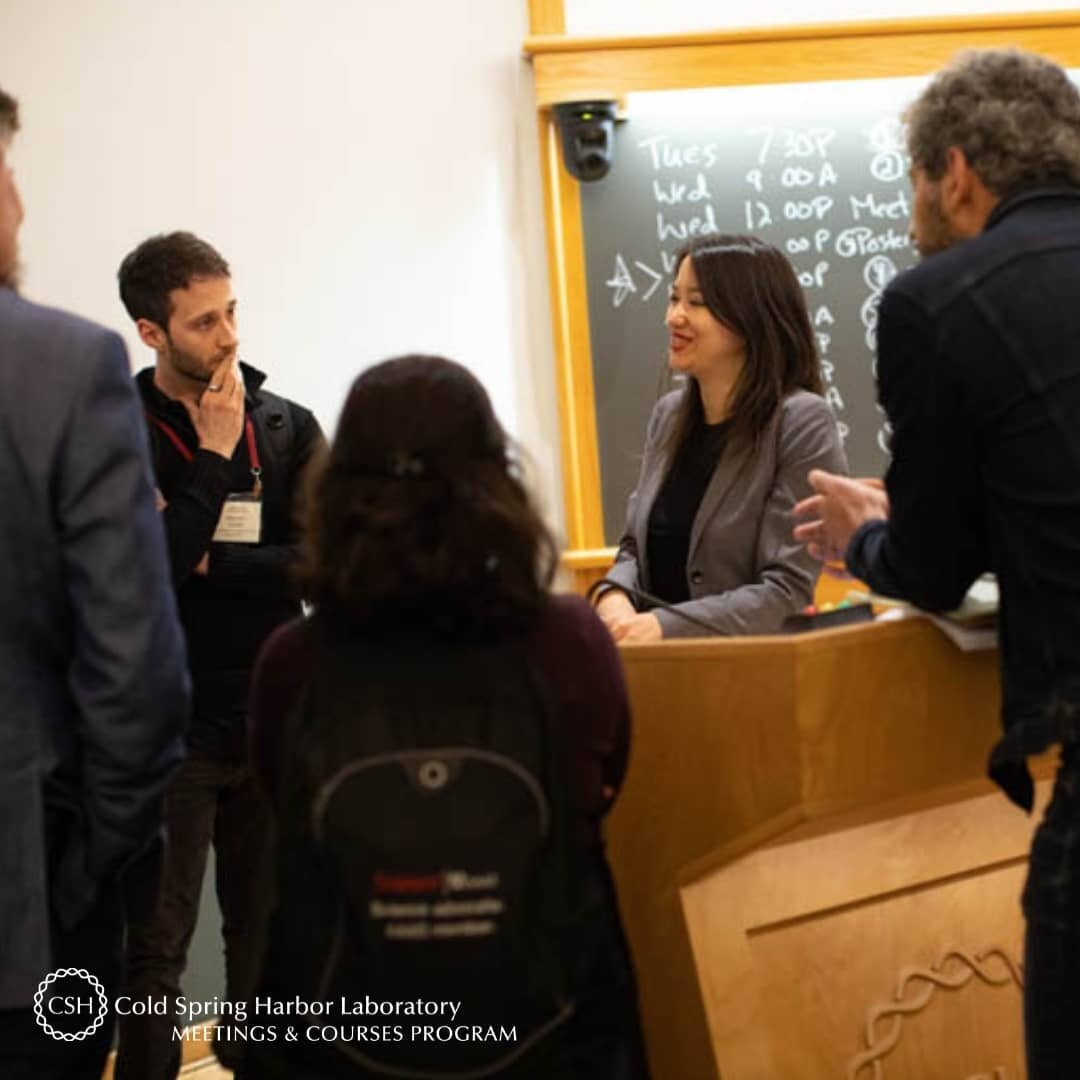
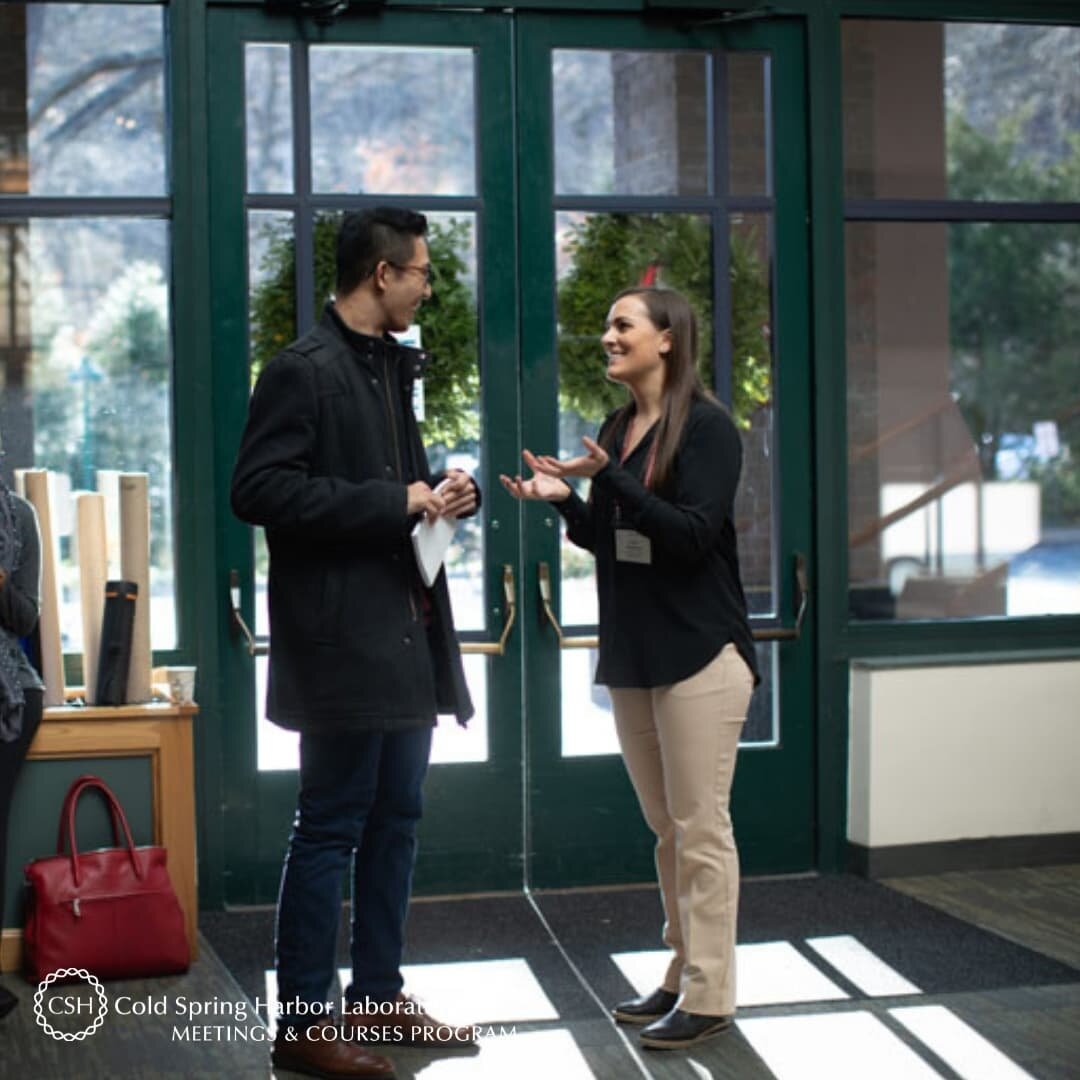
Current Exchange is a blog published by the Meetings & Courses program of Cold Spring Harbor Laboratory. It communicates the science and stories that arise from all of our programs, and is also a forum for discussions on a number of non-science topics that are covered in our meetings and courses. Scroll down to explore our program and meet some of the ~9,000 international scientists that take part in it every year.
Follow us on Twitter!
Are you a CSHL course alumni?
RECENT TALKS
Alice Telesnitsky
University of Michigan
2021 Retroviruses (Virtual)
@cshlmeetings ON INSTAGRAM